
Illustration of chain-forming bacteria

Illustration of chain-forming bacteria
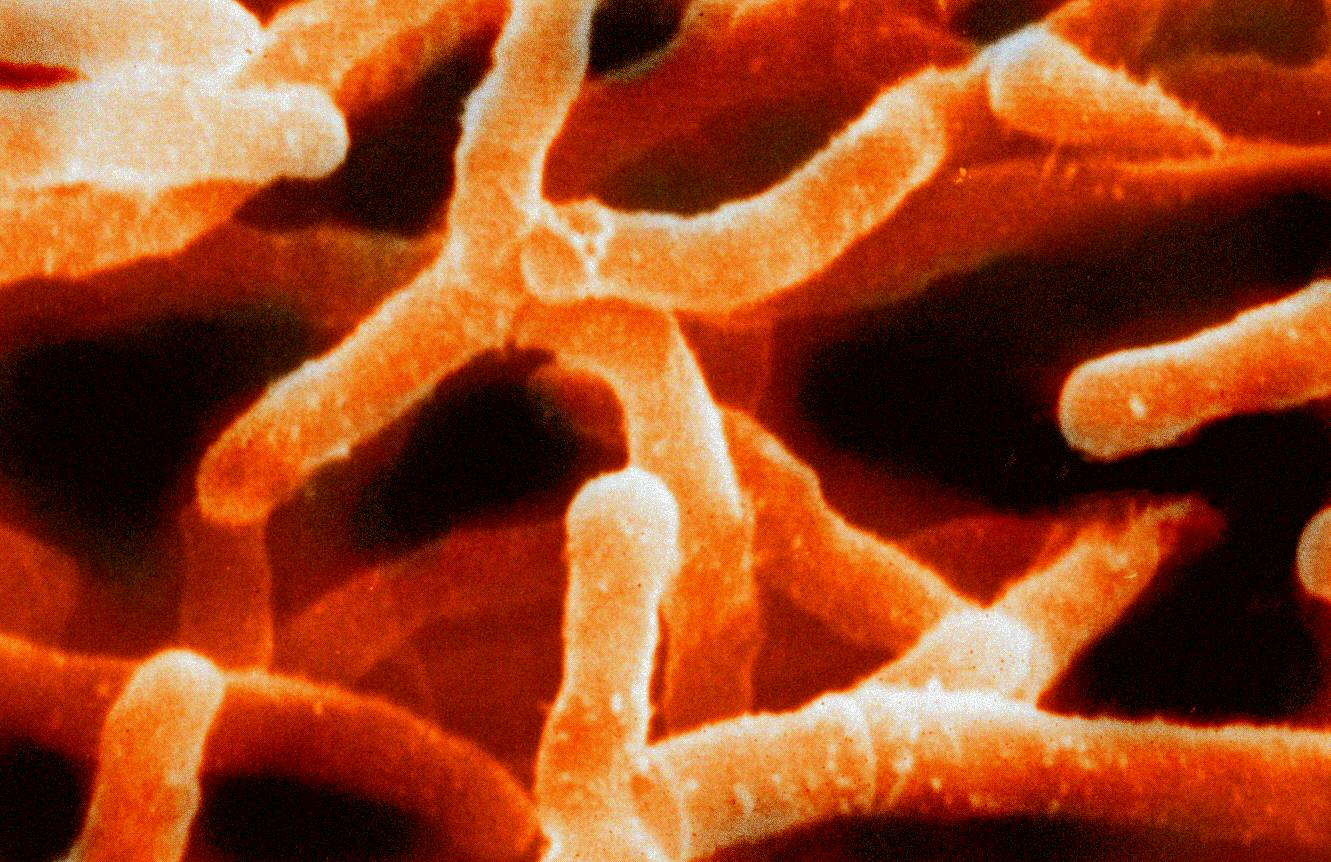
Scanning electron micrograph of Actinomyces israelii (false colour).
By Graham Beards at English Wikipedia, CC BY 3.0,
https://commons.wikimedia.org/w/index.php?curid=4197579

Streptomyces griseus. Scale bar (lower right corner) is 5 µm.

Microscope image of the external filter.
Shows Streptomyces griseus (in green) on top of the filter mesh (in black).
The filter mesh size is 40 µm.
Note how this filamentous bacteria is too large to pass though the filter mesh.